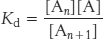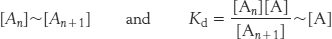35.2 Myosins Move Along Actin Filaments
Myosins, kinesins, and dyneins move by cycling between states with different affinities for the long, polymeric macromolecules that serve as tracks along which they move. For myosin, the molecular track is a polymeric form of actin, a 42-kDa protein that is among the most abundant in cells, typically accounting for as much as 10% of the protein in eukaryotic cells. We begin with a general discussion of the polymeric structure of actin and its assembly. We then examine the interactions between myosin and actin, including both structure and the dynamic interactions between these two proteins. Finally, we turn to the structure of muscle and the roles of myosin and actin in muscle contraction.
Actin is a polar, self-assembling, dynamic polymer
The structure of the actin monomer has been determined to atomic resolution by x-ray crystallography and has been used to interpret the structure of actin filaments, already somewhat understood through electron microscopy studies at lower resolution. Each actin monomer comprises four domains (Figure 35.12). These domains come together to surround a bound nucleotide, either ATP or ADP. The ATP form can be converted into the ADP form by hydrolysis.
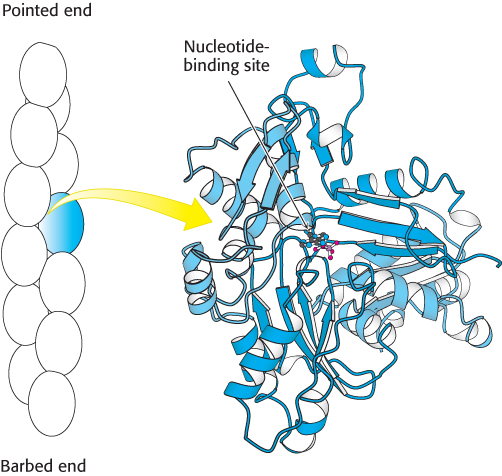
Figure 35.12:  Actin structure. (Left) Schematic view of actin monomers (one in blue) of an actin filament. (Right) The domains in the four-domain structure of an actin monomer are identified by different shades of blue. Notice the nucleotide-binding site at the center of the structure.
Actin structure. (Left) Schematic view of actin monomers (one in blue) of an actin filament. (Right) The domains in the four-domain structure of an actin monomer are identified by different shades of blue. Notice the nucleotide-binding site at the center of the structure.
[Drawn from 1J6Z.pdb.]
Actin monomers (often called G-actin for globular) come together to form actin filaments (often called F-actin; see Figure 35.12). F-actin has a helical structure; each monomer is related to the preceding one by a translation of 27.5 Å and a rotation of 166 degrees around the helical axis. Because the rotation is nearly 180 degrees, F-actin resembles a two-stranded cable. Note that each actin monomer is oriented in the same direction along the F-actin filament, and so the structure is polar, with discernibly different ends. One end is called the barbed (plus) end, and the other is called the pointed (minus) end. The names “barbed” and “pointed” refer to the appearance of an actin filament when myosin S1 fragments are bound to it.
How are actin filaments formed? Like many biological structures, actin filaments self-assemble; that is, under appropriate conditions, actin monomers will come together to form well-structured, polar filaments. The aggregation of the first two or three monomers to form a filament is highly unfavorable. Thus, specialized protein complexes, including one called Arp2/3, serve as nuclei for actin assembly in cells. Once such a filament nucleus exists, the addition of subunits is more favorable. Let us consider the polymerization reaction in more detail. We designate an actin filament with n subunits An. This filament can bind an additional actin monomer, A, to form An+1.
The dissociation constant, Kd, for this reaction, defines the monomer concentrations at which the polymerization reaction will take place, because the concentration of polymers of length n + 1 will be essentially equal to that for polymers of length n. Thus,
In other words, the polymerization reaction will proceed until the monomer concentration is reduced to the value of Kd. If the monomer concentration is below the value of Kd, the polymerization reaction will not proceed at all; indeed, existing filaments will depolymerize until the monomer concentration reaches the value of Kd. Because of these phenomena, Kd is referred to as the critical concentration for the polymer. Recall that actin contains a nucleotide-binding site that can contain either ATP or ADP. The critical concentration for the actin–ATP complex is approximately 20-fold lower than that for the actin–ADP complex; actin–ATP polymerizes more readily than does actin–ADP.
Actin filaments inside cells are highly dynamic structures that are continually gaining and losing monomers. Nucleation by complexes such as Arp2/3 can initiate the polymerization of actin–ATP. In contrast, the hydrolysis of bound ATP to ADP favors actin depolymerization. This reaction acts as a timer to make actin filaments kinetically unstable as ATP hydrolyzes over time, making the filaments more unstable. Proteins that bind actin monomers or promote the severing of actin filaments also play roles. Polymerization reactions can exert force, pushing or pulling on cell membranes. Regulated actin polymerization is central to the changes in cell shape associated with cell motility in amebae as well as in human cells such as macrophages.
Myosin head domains bind to actin filaments
It has not been possible to determine the in vivo structure of a complex between actin and myosin at sufficiently high resolution to discern molecular details. However, treatment of actin filaments with myosin S1 fragments in the absence of ATP results in a complex referred to as decorated actin for which the structure has been determined by cryoelectron microscopy to a resolution of 13 Å. Although a structure at this resolution alone would not be adequate to observe molecular details, superimposition of the high-resolution structures of actin monomers and the myosin S1 fragment on the structure of decorated actin can be a source of insight into the details of its structure (Figure 35.13). The myosin head domain is in a conformation close to that observed for the nucleotide-free form. This structure also reveals the interaction surfaces between myosin and actin. The modeling suggests that the myosin head-domain conformation changes somewhat to increase its interaction with the actin filament. These conformational changes result in a slight opening of the nucleotide-binding site in myosin. This observation has implications for the mechanism by which myosin moves along actin filaments.
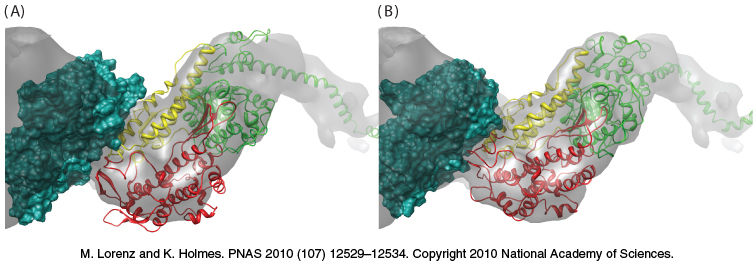
Figure 35.13: The structure of myosin bound to actin. (A) The gray surface represents the structure observed by cryoelectron microscopy, with the green space-filling model representing one actin subunit. The ribbon diagram shows the structure of the S1 fragment of myosin docked into the cryoelectron microscopic structure. Notice that some of the myosin structure lies outside the gray surface. (B) The structure after the myosin S1 fragment has been allowed to adjust to more closely match the structure observed by cryoelectron microscopy. Notice that the myosin structure now more closely matches the gray surface.
Motions of single motor proteins can be directly observed
Now that we understand the conformational changes behind myosin’s action, we can explore how myosin “walks” along its actin track. Studies of single myosin molecules moving relative to actin filaments have been sources of deep insight into the mechanisms underlying muscle contraction and other complex processes. A powerful tool for these studies, called an optical trap, relies on highly focused laser beams (Figure 35.14). Small beads can be caught in these traps and held in place in solution.
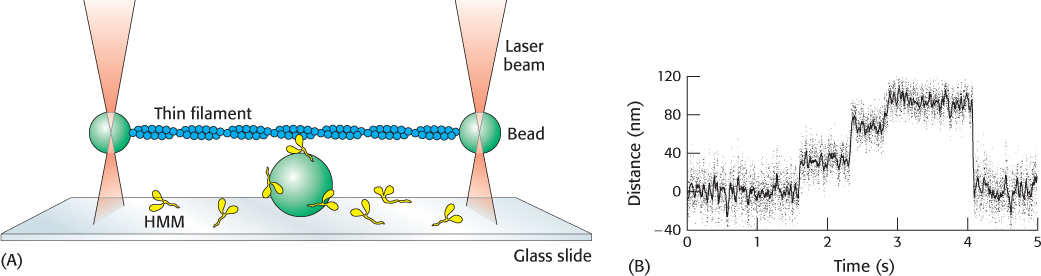
Figure 35.14: Watching a single motor protein in action. (A) An actin filament (blue) is placed above a heavy meromyosin (HMM) fragment (yellow) that projects from a bead on a glass slide. A bead attached to each end of the actin filament is held in an optical trap produced by a focused, intense infrared laser beam (orange). The position of these beads can be measured with nanometer precision. (B) Recording of the displacement of an actin filament due to a myosin derivative attached to a bead, influenced by the addition of ATP. Note the fairly uniform step sizes that are observed.
[(A) Information from J. T. Finer, R. M. Simmons, and J. A. Spudich, Nature 368:113–119, 1994. (B) Data from R. S. Rock, M. Rief, A. D. Metra, and J. A. Spudich, Methods 22:378–381, 2000.]
The position of the beads can be monitored with nanometer precision. James Spudich and his coworkers designed an experimental arrangement consisting of an actin filament that had a bead attached to each end. Each bead could be caught in an optical trap (one at each end of the filament) and the actin filament could be pulled taut over a microscope slide containing other beads that had been coated with fragments of myosin such as the heavy meromyosin fragment (Figure 35.14). On the addition of ATP, transient displacements of the actin filament were observed along its long axis. The size of the displacement steps was fairly uniform with an average size of 11 nm (110 Å).
The results of these studies, performed in the presence of varying concentrations of ATP, are interpreted as showing that individual myosin heads bind the actin filament and undergo a conformational change (the power stroke) that pulls the actin filament, leading to the displacement of the beads. After a period of time, the myosin head releases the actin, which then snaps back into place.
Phosphate release triggers the myosin power stroke
How does ATP hydrolysis drive the power stroke? A key observation is that the addition of ATP to a complex of myosin and actin results in the dissociation of the complex. Thus, ATP binding and hydrolysis cannot be directly responsible for the power stroke. We can combine this fact with the structural observations described earlier to construct a mechanism for the motion of myosin along actin (Figure 35.15). Let us begin with nucleotide-free myosin bound to actin. The binding of ATP to actin results in the dissociation of myosin from actin. With ATP bound and free of actin, the myosin domain can undergo the conformational change associated with the formation of the transition state for ATP hydrolysis. This conformational change results in the reorientation of the lever arm. In this form, the myosin head can dock onto the actin filament; phosphate is released with concomitant motion of the lever arm. This conformational change represents the power stroke and moves the body of the myosin molecule relative to the actin filament by approximately 110 Å. The release of ADP completes the cycle.
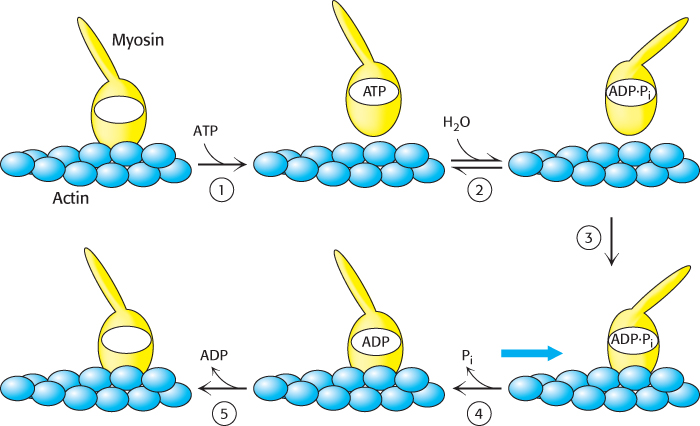
Figure 35.15: Myosin motion along actin. A myosin head (yellow) in the apo form is bound to an actin filament (blue). The binding of ATP (1) results in the release of myosin from actin. The reversible hydrolysis of ATP bound to myosin (2) can result in the reorientation of the lever arm. With ATP hydrolyzed but still bound to actin, myosin can bind actin (3). The release of Pi (4) results in the reorientation of the lever arm and the concomitant motion of actin relative to myosin. The release of ADP (5) completes the cycle.
Muscle is a complex of myosin and actin
The mechanism of moving a single myosin molecule relative to an actin filament explains how muscles contract. Vertebrate muscle that is under voluntary control, such as the biceps and triceps in your upper arm, has a banded (striated) appearance when examined under a light microscope. It consists of multinucleated cells that are bounded by an electrically excitable plasma membrane. A muscle cell contains many parallel myofibrils, each about 1 μm in diameter. The functional unit, called a sarcomere, typically repeats every 2.3 μm (23,000 Å) along the fibril axis in relaxed muscle (Figure 35.16). A dark A band and a light I band alternate regularly. The central region of the A band, termed the H zone, is less dense than the rest of the band. The I band is bisected by a very dense, narrow Z line.
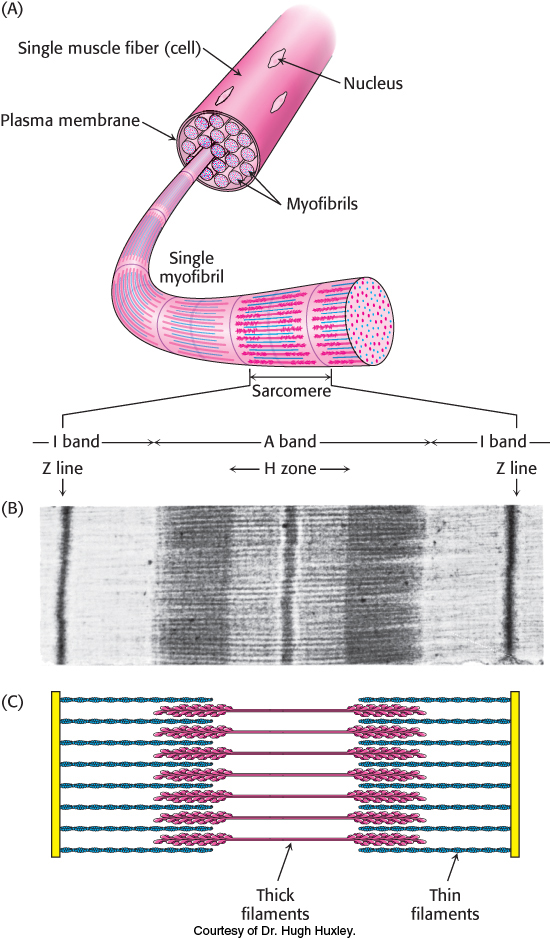
Figure 35.16: Sarcomere. (A) Structure of muscle cell and myofibril containing sarcomeres. (B) Electron micrograph of a longitudinal section of a skeletal-muscle myofibril showing a single sarcomere. (C) Schematic representation of the sarcomere corresponding to the regions in the micrograph.
[(B) Courtesy of Dr. Hugh Huxley.]
The underlying molecular plan of a sarcomere is revealed by cross sections of a myofibril. These cross sections show the presence of two kinds of interacting protein filaments. The thick filaments have diameters of about 15 nm (150 Å) and consist primarily of myosin. The thin filaments have diameters of approximately 8 nm (80 Å) and consist of actin as well as tropomyosin and the troponin complex. Muscle contraction is achieved through the sliding of the thin filaments along the length of the thick filaments, driven by the hydrolysis of ATP (Figure 35.17).

Figure 35.17: Sliding-filament model. Muscle contraction depends on the motion of thin filaments (blue) relative to thick filaments (red).
[Information from H. E. Huxley. The mechanism of muscular contraction. Copyright © 1965 by Scientific American, Inc. All rights reserved.]
To form the thick filaments, myosin molecules self-assemble into thick bipolar structures, with the myosin heads protruding at both ends of a bare region in the center (Figure 35.18A). Approximately 500 head domains line the surface of each thick filament. Each head-rich region associates with two actin filaments, one on each side of the myosin molecules (Figure 35.18B). The interaction of individual myosin heads with actin units creates the sliding force that gives rise to muscle contraction.
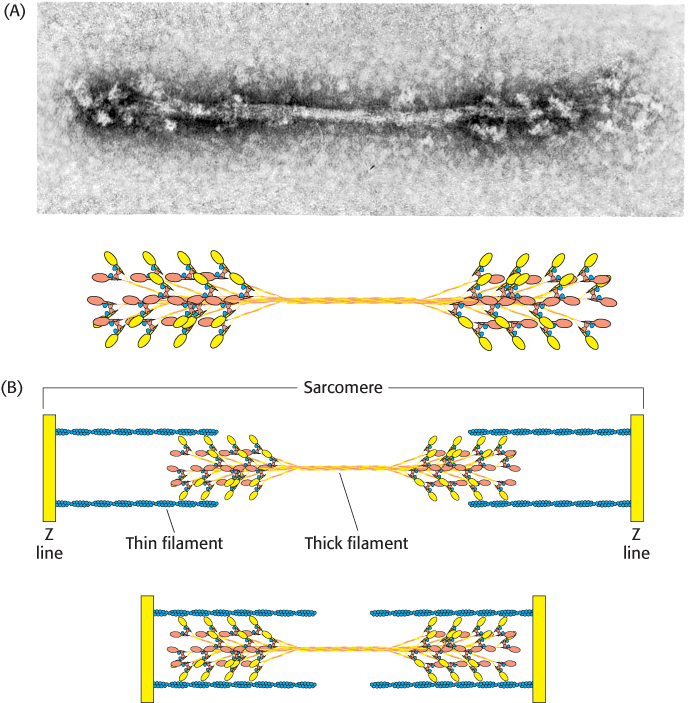
Figure 35.18: Thick filament. (A) An electron micrograph of a reconstituted thick filament reveals the presence of myosin head domains at each end and a relatively narrow central region. A schematic view below shows how myosin molecules come together to form the thick filament. (B) A diagram showing the interaction of thick and thin filaments in skeletal-muscle contraction.
[(A, top) Courtesy of Dr. Hugh Huxley.]
Tropomyosin and the troponin complex regulate this sliding in response to nerve impulses. Under resting conditions, tropomyosin blocks the intimate interaction between myosin and actin. A nerve impulse leads to an increase in calcium ion concentration within the muscle cell. A component of the troponin complex senses the increase in Ca2+ and, in response, relieves the inhibition of myosin–actin interactions by tropomyosin.
How does the myosin reaction cycle apply to muscle contraction? Recall that hundreds of head domains project from the ends of each thick filament. The head domains are paired in myosin dimers, but the two heads within each dimer act independently. Actin filaments associate with each head-rich region, with the barbed ends of actin toward the Z line. In the presence of normal levels of ATP, most of the myosin heads are detached from actin. Each head can independently hydrolyze ATP, bind to actin, release Pi, and undergo its power stroke. Because few other heads are attached, the actin filament is relatively free to slide. Each head cycles approximately five times per second with a movement of 110 Å per cycle. However, when hundreds of heads are interacting with the same actin filament, the overall rate of movement of myosin relative to the actin filament may reach 80,000 Å per second, allowing a sarcomere to contract from its fully relaxed to its fully contracted form rapidly. Having many myosin heads briefly and independently attaching and moving an actin filament allows for much greater speed than could be achieved by a single motor protein.
The length of the lever arm determines motor velocity
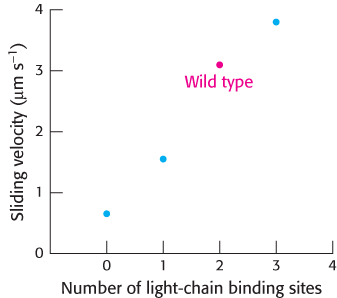
Figure 35.19: Myosin lever-arm length. Examination of the rates of actin movement supported by a set of myosin mutants with different numbers of light-chain binding sites revealed a linear relation; the greater the number of light-chain binding sites (and, hence, the longer the lever arm), the faster the sliding velocity.
[Data from T. Q. P. Uyeda, P. D. Abramson, and J. A. Spudich, Proc. Natl. Acad. Sci. U. S. A. 93:4459–4464, 1996.]
A key feature of myosin motors is the role of the lever arm as an amplifier. The lever arm amplifies small structural changes at the nucleotide-binding site to achieve the 110-Å movement along the actin filament that takes place in each ATP hydrolysis cycle. A strong prediction of the mechanism proposed for the movement of myosin along actin is that the length traveled per cycle should depend on the length of this lever arm. Thus, the length of the lever arm should influence the overall rate at which actin moves relative to a collection of myosin heads.
This prediction was tested with the use of mutated forms of myosin with lever arms of different lengths. The lever arm in muscle myosin includes binding sites for two light chains (Section 35.1). Thus, investigators shortened the lever arm by deleting the sequences that correspond to one or both of these binding sites. They then examined the rates at which actin filaments were transported along collections of these mutated myosins (Figure 35.19). As predicted, the rate decreased as the lever arm was shortened. A mutated form of myosin with an unusually long lever arm was generated by inserting 23 amino acids corresponding to the binding site for an additional regulatory light chain. Remarkably, this form was found to support actin movement that was faster than that of the wild-type protein. These results strongly support the proposed role of the lever arm in contributing to myosin motor activity.

 Actin structure. (Left) Schematic view of actin monomers (one in blue) of an actin filament. (Right) The domains in the four-
Actin structure. (Left) Schematic view of actin monomers (one in blue) of an actin filament. (Right) The domains in the four-