Chapter 1. Chapter 20 Branched Tutorial: Restriction Mapping
1.1 Problem Statement
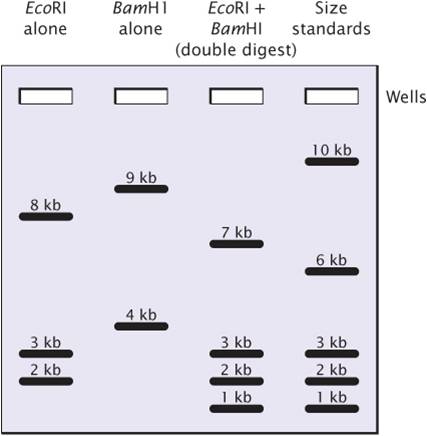
Restriction sites can be mapped by comparing DNA fragments produced by digestion with restriction enzymes used alone and in various combinations.
The experiment began with one sample of a linear 13,000-bp (13-kb) DNA fragment being cut with the restriction enzyme EcoRI; a second sample of the same DNA was cut with BamHI; and a third sample was cut with both EcoRI and BamHI together.
The figure at the right represents the separation of DNA fragments by gel electrophoresis.
Question
Provide the correct restriction map of the linear DNA fragment based on the results shown on the gel image by choosing the correct enzyme names and fragment sizes from the associated dropdown menus.
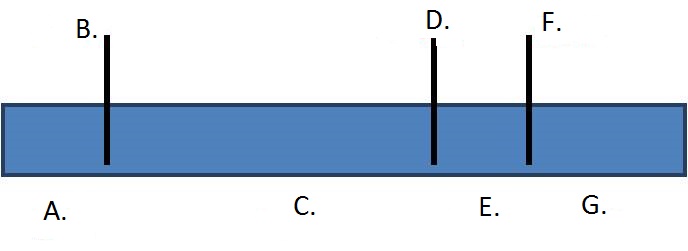
| A. jAW2nq2QxNaPmtooReC4oNUUyKG/fW3ybjmW74MKvUE6jD84WLaNITJx8ki/7SZi1Gvca4iV4dxkOG38GKeoToo58tR4RVN433Ns/it5yWo= | B. 4IFBf9ScuaDHv4T88pKVLY5cyOOOx0XKqZbHdQHyd2kPNeFTaACHltwZ7ojhpdwQUSPOWWdaiHx9rq6xLJEZUqToMduwogkP9Dmsd57wJTk= | C. TN8pRmL1WdrhZ2x57C7AmhEk+u50Sg/Tak0OuZzK+DWIJwaICEahWdviPD9p5FPXXwe+KeU3yja6aqx0q9P7VsSjnn3l1cLmdDhRf9Afwk0= | D. p8CSlrXINuswoOcdDxwvC9HTT5XW6BVO047taLVdelAuzYN49m3UReW0PPI5CpB+xLnsTw8O8x8Kyqxofd3v69FTcOdbs7nIMSzpCSnEKRI= | E. pWZZxi3nhJchInvjqYiiq2rCwh/dPk0r6zgH4BX2Fh7YSmnpXPJmXxP3SXF+qdyszYFd31aCw2SFY5m6PZ2kYeDgLAwVszy1bgJC3bOdD0s= | F. 4IFBf9ScuaDHv4T88pKVLY5cyOOOx0XKqZbHdQHyd2kPNeFTaACHltwZ7ojhpdwQUSPOWWdaiHx9rq6xLJEZUqToMduwogkP9Dmsd57wJTk= | G. bxslCdwuwP4Bj31zlKqGeC4guY+f6ohY42FeTaqmETT3oTntx+8DTcWd+pgGrs2LDIZcpgQbYja//ttI42YlKj3BlVHPl9f1Zc80Mr82YUA= |
1.2 Problem Strategy
Question
Determine the best approach to solve this problem by ordering the following steps:
1. Examine the results shown on the gel and compare the single digests with the double digest.
2. Identify the question that is being asked and focus on key pieces of information.
3. Examine the results shown on the gel and analyze each lane independently by constructing a restriction map for each single digest.
4. Construct a restriction map based on the double digest.
Instant TA
Begin with the basic information and individual analyses and then build upon this information to solve the problem. The best approach is to analyze each single digest independently and then construct the restriction map of the double digest.
1.3 Step 1: Understanding the problem
Question
+Prcgcf3Xd3ZOkGWH0XuK+Ymi5POiQUmH9J/6hYh9XYVTQm4gMGQSWhm3EOtaRWZTqYSY0PPqYfbW5leXSWMO1AA+FCfw3SpiHrZqMloluEob3UmdhT+4WvMWPyFMhPoA0J/FNkknf+U8eZtSsvzfQxZVSHFUcTw0XvRkZYyA+Ki4HQWccaWvSBMMDqcjfvy6S1uxuG/GTsVo4qLwx0D7tHF0OFEeOuRo2vOa9GIbUruCKiCw8Hdh+UnK+V9Tr6unJUpa+GNZz0j4U1XxtrfT5CkTeBfmGgt1leCz617SRF5zOy6Epv8FNHWYFw5YspVqHFY2NUu8Ja8Gwi1ma5kNV1JUYru00Ygvc0ZfuI1f07pBpDTzcpKwt7Q8Jn+Llh/LmCjmxFJpf/aWNXzbQ0xTaSxrk6c92CI15Fa/Q==Instant TA
DNA molecules can be linear or circular. Read the question carefully to learn what type of fragment is being digested.
Question
5HhZJ05Wif8FBnjS9vBPzXQ39ZPfCifdOhcKZjvLSSWW0MINQD2dm/i4y2rFKzHdGtxjfr3iuHcB2ftY/4SbzQOzmpdz/KaZaGGle/r98/buo1ziVAQJd95GFYQMSz1/rmAiYFUJ5GEy8npRFMBY/wHRy6CYIHd07yDVt0aNOg8a1de4FVfayz8nOSMUioWBpLMtDQ2tdnf4G+sR8GByIdWNkJdAgFetK838eME0kUNjUX1DjaeB3AqMCzHHu+GRTTFJTRxsmyksYxaHdI9gNbW1aN9ieMMDlsHUuDOo7RHwOoyNFbLHm/2GO7GtmR7rDQMVoluzdHXV6h8Hit58TrsqpfYqVGWYWLdRZo2f4bkafsxkGNGxXCgy8odiVKwo4/8e1qREAL/uDa5aAhzNjUeMXujQDUtgHsJ0c7RyddC1N5Y/G2+OgAobuCKWvD4hG+vDbvG0qkSFxDOcHKkuNbML2jKzl2IkDxGj6i3b2TPG7a9cwvzj2AXqH03JQMW0DPkV6sS6g6tfIjHdt98usCBWP17sAjoTgfiVK0OrPpce2fA4cAEm8G/JULZe9pXzNMHDVJ7juXeFaTmxJEae1Uf5BXVgVginH/GDhQ5yJjN4K7WaXbCTTs8McKd4DypJ5NACAaY8VYDlY/M7pXCN6UgVkBOkg/1WgpvC9ObEcjtuqhofKnXVO1+NcyVkn66XG/mf/uTH+Pdvu63peJi1qc9OmowxGyTA32glqo7Qt86DDLXasAwWzoP6M9RQ/x2RIJwsUT5ksl1dMVQzuoGajYPNbKushFmjRc1T8H0B0ksG9tVcXczU5Xka9EgrdLQCW6llLlTrXQ5mjGwCfluB6sB/4FKBBh2rhlzFTChdJMKAn+eKZpA9ZE+c6SYpejCqZKK5IpMlTOkJYhJqhNLHqLwNnsBSCIX4PJbv5Bhqy5izC8oFh1bSx4GVLR9W2DIbxmnW3ApCyuQLOi7dFQZrT0ygFnLmPDYqPYT19srZtTMyTbiYTkApiwfyNNURdDDTq/kYZw==Instant TA
It is important to know the exact size of your starting DNA molecule. The fragments of each sample lane should total to the starting size of the DNA molecule. Remember, the size standard is not an “experimental sample” lane.
Question
M9ynbxY5VIjfONuGAuzfEx7fckcfdAc7kufIshc/JcoSHbnVLcZlzM5GPVNUNrqot29IL2TxjV0kR1wxb2PDwI0yQbnO0FqCbno/dAgTPxTDNJKeX7HnFhI1qNelRNnLcnR21T2w8jGEIi67XuU/Df3lJP+F+e5pQ3WbTL/vPaQYcllarImrs0Ss6NtfjEdphCK63RiurSJFW3zChkHfRBTwxvew5oxSrBNOvWo560tO1ti1EovufT2N4Kz1x880WAhv2DtMIlo9llhfucBWU9Qs6gad0xm4kpsPaofkjrd2XKBkh7V7MvrUrnsFLyEgl57QuJKsG8QjJkUjjby9KMf9iPqNcQohThsTT3B4RqB/lsjmV+nhMw8fZ7fELc2Uxl8bvLH4rpzhbHxsrP1me531fzvdHs1TPxK7RPvoQkS+38mMeriDRvuhmFMQMdHgZ+tFD6VgU5Xw8SpdGS3BuX/q+WG0xlQMoysKq0PGs1TAw5jFKze+38u7oDy8E4VpojTyMfwXnUZEg2CcHo1r7Iq8MhbIjxzjjSCeDlJM4n5JRNi507q/0ajzhJ7cz9USnnJiXAWA7F9bmefRJzwUGpfMbRPP1lYN47rrva3Uc6fTm9CaKQS9DkGbeHCGEa84RYEsbYmAREo=Instant TA
Restriction enzyme names are generally 3 italicized letters followed by a capital letter (normal font) and a Roman numeral.
More About Restriction Enzymes
Restriction enzymes identify specific nucleotide sequences in the DNA molecule. When the enzyme binds to the specific recognized site, it will cut the DNA molecule. You can find the nucleotide recognition sites for EcoRI and BamHI at New England BioLabs (just search for the enzyme names).
Question
iZd+gl+GhL9/gyjOZ3G7bsq2n2aT5yXoouJ0h0m+21Bac8D95CpT8pvpy4QttoJ3+mDlqcFkMUKVJDPqBazBjVyCQl2A8OATxFd2tuNNRVoPcQ2qLkZhhGbArCMAtFliUy9zQ+P52LGApAA36zb9wviQadwjRkp8LLAR/BzkhS0fYLvI3JFb4gvuIf+705IaMEigCGf1RtGejfNPz4g4pLNA3ChQ+x3UvGgc6JfVeErQ2fc62rm5Os8S/t2UpQ9NGDI6Xd6FTRjDTDYComwE08QP5O4nvZK8l7e7au7WjGI76JLOKbtxbA4+BW6heG0eZTkfCCD7C667lt6EojEDjwk+RIc7XDs9V5/IhOww/B3HI2kUUyB2l8Ev8uwOM2l68OM60Kylyii5U5LC1y0Ma6Eqi7TjXbeilcfU9gWM0SIqlz3dA1lwH5ARAuEgnaLV2Vj3gCAFriyeMFCIAKW19JT89ZuCFH/CfliSaWZ+71LEPJWrGLSexmevCo0LR5vMSK7hs9dDkwXvAAuCpB7CSfif4bnk/2VX1vf05Gv3GO1eZWD9gFqtFzdRFckuk3SoZ7mipU1pangqKkvmPGmaOxzQNEe0D/cayb4fEpxb48b0fAzbkm3OcFkGZOoTomzSnbUN3sPgWfHX3AUSYyxJChtkpB5khsjGl29V1qx9RPEnpgLF+fr/M5/SlS1Wdcw63MI3hy7d4zz61xp1Q6OY7iwPzhoKY0xLXOOKb9zYYNmn3bBT48Q+HIRmo8cBIAgrpWL4ayQ68qGsxojlzYNwmCppTIsdbUs9ziyEBrbBs0cfY7MIct/d7fv50s+kGuvDKLcUzvPe8j/e82v1r/5bkrPfVEZtUqZZ5JIPSwuOp0WLnjeJsLHm/c4mUf999jc69HEtTRM+SvancfaQOh/CPXVW3w4VytxPxi3skPyLhEAofK91D5JoWg==Instant TA
The restriction digest process occurs in a reaction tube and the resulting fragment sizes are mixed together and undetectable by the eye. The gel electrophoresis process separates the fragments and allows for visualization of the individual fragments so that a number and size determination can be made.
1.4 Step 2: Analyzing the Gel
Question
acJ6+sOePgH2tnTcdNcLmwrUxDnQl5cUBFiB63Vqbrq/72bVwybtJL585Moc+2U3QldjgnQJQE55L6DvoseQiRxYY124PtiXRE4ZnWqpn8gSaC8rb944iiTVFp+9wGAZ+3smSbtIbPED64WpAxkFiiB4ztVhwyGcMAis1dV0VzsUTUagSEnKXlJhJ0dGOq5V/b5shsOmgJp7xLJed9pt8NiInNedwxUAaMKF6UjS1rke9MRUc3Eg+Ic7nPxWsTaRiVviJANH4ayiV/xhX4nE91xkbnEi5pgDAcuMsG8X15jwos1LtYGhSJfEG6PHB94Pyfj5YmCB1mAstVmzYgPVAvzoRYdvF0cBxxOZD0VCK3PYrcKz4u5QW7xSIX72Kpt8Qr1uNpz9DKHBY6cs2Y/8QwvZTYEXVoJ4gyI9K/TopWfkLsz8+5HNNrTVw8r6Qqq1InrP87u5T4PxHgjxfNVMpOTOpB0oXLYaWrnSca4QC0JJUEYD3MMBIjlQRZecTgEFtu07VxbOZA4ajEf3t+jtDOryRvKXyZ+e28rIsiLP8cXdnzNzplzrI+hCHDDm9g2Rp6+V1EA0iAap/FOUgY5a6p5faFP2bVyIomuKXhI9mMA=Instant TA
Focus on only the lane that is labeled “EcoRI alone” in order to determine your answer. A fragment is displayed as a band or single line on the gel.
Question
DsYOlWy4eszx95aF2f55mXo92JIRvXhwJgGs6PAHJLng0NNjMdKUWIdh7v66ctRG4rkdl98mPmGq6IFcicwQoK9aJ2WUWcV7PI+QNqRWNdw5KuCtSwF2YNEq5EhhWUZv7sCn1dpPZrZDONjPhCbvEFrAl2Hl0iJJKprp+nS/eeexFr+nqGWD/hEHMSCpPAOAL1qk3/FCSJ7xvgz74xmGmUOEIoOLQEgx8Rdx063LsNMzQuzx2PTx/pz2kfroS2j+bSB2iAL1EyxULubj6Z7/Gj2K4eFTrtT4Pi4KoqjvJ9KSLH/Jphx9KVhg+m0RY0/cpEN4JGVvzaPoAhG0qBWFhJX6z6cG/2K6yRt+JCwsGIrCe4MHapEuOLwzpuzC37zJZyHs5si3cNqVG3FeXYAAtw==Instant TA
Think about your fragment as a piece of paper that could be cut with scissors—for a linear molecule (your piece of paper), a single enzyme recognition site (cut with your scissors) would result in two fragments (or two pieces of paper).
Question
3. Draw a restriction map based on the first lane of the gel. Correctly label each restriction enzyme recognition site and fragment size using the associated dropdown menus.
Note: there are multiple maps that would provide acceptable answers for this problem. Use the relative sizes of the fragments to guide your selection of the correct answer below.
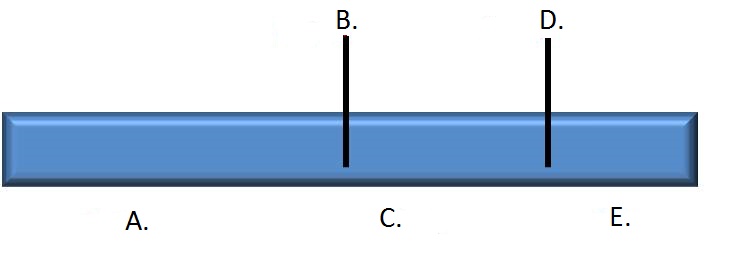
| A. pyRPnJLljT/bv5bNIRoYuJgIx/lDd0szQfb5ocmXqJsygWnIZrTPNHFBDZvZCbtCrHUz95A63s2kQXVnR3+Ghr3etMrE+cy4h68cEuBjkwk= | B. 4IFBf9ScuaDHv4T88pKVLY5cyOOOx0XKqZbHdQHyd2kPNeFTaACHltwZ7ojhpdwQUSPOWWdaiHx9rq6xLJEZUqToMduwogkP9Dmsd57wJTk= | C. 2cBKgkaMWxCdupPbnwKXk+VviUJwyw1NJtIPOWO1EQfhx23BhXv/R1Y29RPUsE7IhLqDdzmJBLLHN6bRmspklvMF/RTmYDYzJyygON9MN2Q= | D. 4IFBf9ScuaDHv4T88pKVLY5cyOOOx0XKqZbHdQHyd2kPNeFTaACHltwZ7ojhpdwQUSPOWWdaiHx9rq6xLJEZUqToMduwogkP9Dmsd57wJTk= | E. nmfXUjvCRtsQizKjuQKRE1U1zMzZdAfQac7CsgJ8jCpZhL89mOYNixkFxHIzZ9KaNPrxWwk1lXQ6Ojmiqkh3HIpRkkzEUguxR0+JApMipaI= |
Instant TA
Focus on only the first lane to find the information needed to answer this question.
Question
I2wxS6GBgabAuoqNmQfrFLiyk3wxzdqem18IheM9+by2jkFrm8+QU8UtdjCELNj5tqkRwJmNWJEQbArHUXEiZmCqhvrdLmsoFX4Q74Y1dD5GgsXqUwLYLSHp5JWfpzSTFCJbuXC+kaYIqMil1GbWybjYUPdxO3HNz4EkoVw3tx7om+TrSc4NqAFzyo/LujD04iTX1Ui3EGWAApAO7/0M3LL7dnzKb/JvRorkWZuNyU7/bH4LU6ZWbLw2kTSKkCYCnO4TP8NxklNCtt/Kr1vETz+4r1BeEFgaDq9CZtDPY2G9aRNDkVPcrYckTt5Stwu4wa+fTBby9iCYSoPmz6ih47HmajExm32I2dU9qdYpASJZfgMlIaMphvu3yM+20hb5QIiSP0qYWxO5TWCcCJyQIopHcF/5udT97LcvS9EHluDhcW6d7gvw/95Z2xj/oZ31xWInws9JebWWgNCv7gNRj4NBsTMpZCRaI5ZW1W88moGDp35taQ+WXo5iPvTVMmWT52+4S5EPUl55AWVU2vvIZs384uFInKUAFimk4eh2CUldu20HSewA7ARi9lOCs0v2EZvG8ZlTFbzLqsR1TN9Bi0KHB/B1BfhEjEI06ggpWVA=Instant TA
Focus on only the lane that is labeled “BamHI alone” in order to determine your answer. A fragment is displayed as a band or single line on the gel.
Question
h380Z2lI031n+0oI8dv4MJiCBNQH4f6fGmqYnZwkj1ELlLYqHwM4MYcF7SgopQOUICkiUMdWO08iPXzYEuFAdddMEY+8dLjdh9jfYkYKmCUfqkuxv5vJ3DBiHy3k+zNqjEyXVefXbslhWjHjRFB4L8Gjir6QHR4JIYokr4OLjPsIGbO0+N/9wgTTa5pMk4uwgr7gVxQ2HRnN6dm+NNkeUXhj78fWSZt26EgOrr5KHImx7GwppVq8BcaBfSu9L4l5CwagmUCxg6utaz/1ckRR0Q1GpPaII3vhBkCFDVA6Xxrd4kDtQrVZ3o6HCWC8UUGANMT9ZoHAeu+ik4K3eF3Auvq/INU3nh6BRk4/kbf14bOAAIziDG5FD2DY//Ek6KHJv++7BwofVSpr/7mOInstant TA
Think about your fragment as a piece of paper that could be cut with scissors—for a linear molecule (your piece of paper), a single enzyme recognition site (cut with your scissors) would result in two fragments (or two pieces of paper).
Question
6. Draw a restriction map based on the second lane of the gel. Choose from the dropdown menus below to correctly label each restriction enzyme recognition site and fragment size.
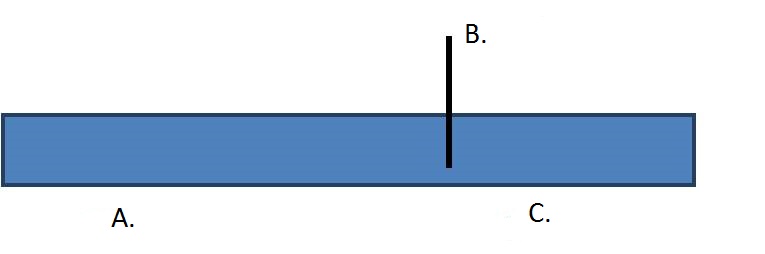
| A. lHkCjqHWlhX0HwKkQwrppLIhhWL1PJtHa+9UaPdTu4XzUoXRggy3n/tN6sEePAGC32lXr8U8CrhivexQLk0Vl5IC6UrhqO4enoAQ74sp9r4= | B. p8CSlrXINuswoOcdDxwvC9HTT5XW6BVO047taLVdelAuzYN49m3UReW0PPI5CpB+xLnsTw8O8x8Kyqxofd3v69FTcOdbs7nIMSzpCSnEKRI= | C. a7Te7QcQkYQuzxFDcc9igfxBbz/BU+plMX47YZ4qXwmWnjMZaEaZB8ON7FJ3tYFbWIxyzPhDf81tNZFsWOrav7XRSlAkYjOVlUnStOPOUfg= |
Instant TA
Focus on only the second lane to find the information needed to answer this question.
Question
T6kvQ0dc+BqpXVbsk5PIzh7ouWfanE0JSwixtBh/byl9aH9nb7fY1h9ujM93uA+C1eV/xlpebdLBVYvdwKx3jKIaFuGBkzubdTgB+zvaIc5rmlZ1xhRyaKBH7BT1dCoeuY/3YPGlc19/Y81FDxFXHQyWtIO+VqMN0O0ACBkCgMcgg7T5Z4TPmS8q7sw20i1c3WEoXUNklujz27DoO5k4eO9VePzYffczU7ihW7ZTwFRnZVV9kFlcawxZ+jhu/3IE+Yiz9yEb+hfWFEbPGVYAL6ZJst/6knwNar+OD9Y6E+3KMvNmM7K+NLhgJjEjTNvPlm0bzSrGJQ0MwufvT8qb25q0+/jpM4RNgBCE64l8wtWDcGNd06xTQUFrczZnSeHbIF7NtcP7dNEe6lVg8ENezNnqsA5H7lcSzwreOcAwi/LAQ1G/IOQPg5s15WVUpAtPPqMKj8csfBjqQQ9Kc2aE9dsv79Kr+B2weuuXoEoSVv6fZ8xbWAMMW6Uw+VOw9ddP0oyjit9tECnEo7zsN7SwuiNMmn4GWK/4z6TpxQ5p9W1LmwVXDVdI9Ws8T5RVCGykjSFpFgNbNEScTNidXX3kaFUivgISGO+1pu597pKYtAFsSLkkInstant TA
Focus on only the lane that is labeled “Double Digest” in order to determine your answer. A fragment is displayed as a band or single line on the gel.
1.5 Step 3: Understanding the Data
Question
jBPt2MoKVqiABvqnyxS+4CRnAcCmp14JWpuUh1gKkgmpXlk9g9w4X2F15qA/gUQ8Ev0pFNaktzIjIklVqPvjOm/IsXz6QefK3dm/ecLqsm8fVdaBq3vK9PkMMmPLexhAHALtzrMh59Cy3cZB0rMYR67Id5KlOQnfH9vwHaz0zCfyAVDwoLix9CC7Okp4jJZOwUCObU6ItJDk3XqIhs7yJWG71WSYpesqscJNaa0Ylmf4//j2L8weeEQ1mrzqZ5QEahgG3KA7iEe7Vy8/kbVxGM2vmJ49eMIh8XEhflQ8Z0Bcwr/NYyfzHWUAi5+G/8fkAlQt5b8RNBCS7KvlNFkQ4FCYWzN6R8eW3hLEsS69iZZ5qtd4Xq7KbbhrL8Vaes3Xxb7lsTE3QAhVHLsYm43Izw==Instant TA
Focus on the lanes that are labeled “EcoRI alone” and “Double Digest” in order to determine your answer. A fragment is displayed as a band or single line on the gel.
Question
7wFJBoiFKQoQ4VgMY+fCYd5oykaXrxGz83CFoDskYQ6E+5qD7NemQKs9jIn3W8lbnD9GpzgZIeNeHwtc2Ws0EXTrBIBadYb+wB37yKYzn3xdVeUYl+a0XuigXARWOqhp7VP3DokrMZa8LmmQ8F6E0vaNJZbMZhnkjx0lbY18Df5nBw9hHSm0FWO9GTQj6Qm/zoMTrxQIgmidq+/XeM5x/+EruxniyR1xRJMuqSP7MCNFbYlsOx6RXuB33Mt4n5PBqE5jMJ5pwA0ZVwUYcPaD7zpdHS1RI1Nmy/rFD68CbteuHu65apsdmsAeFpvZEynNZ8u4c1+MSIsQLcBoBiyHyx1SjyQUOypjGpZhFdFAKNWqIZyIyKJ0Rcn3dK/1JkcV9z92OdiZr8ARMpz9QKwfPZZ+IUrTC0ppjNEHuXVKwMyEd58w7w4NAgns47/zesulI5eHm1cHXuwdODkOj8XW/sNJFnUocCI/UpM/uWBW+QfmB5487qtIdsEWXjg37y35cQWXHHRzst8pMbK+/hiIX3tBVCuf14ETx/9W54tPH7VSkzP1/1z9r/wX97Gf4GFy5Wo6hL2SSyDX39rhN3+JscPy8wzSX1ExPVtYWXd7iLSLAL2SwGifkUznC5Y3gn97XRhHeUAnzME8RzKsEmaMoMiEcgqH8bwERNvD3eYnsBisSWViLSOtuinW/edqwJtiqoJeZtqojKjMujpn3OhFFTM9QNRoHj8mc+nVR8pZ5o1H5rDkGYZ+XG89yRdiqjz/oexO+vEXwE81LlM2j4W5qyQPx3rLyp94n0pOyZZrQpSj0QmfcWtcibm10C4+wxNvvBbsCIug7wTqi7zn995UYxCNCTayOyUDInstant TA
In a double digest, there may be some fragments that are generated from the activity of one enzyme and some fragments that are generated by the activity of both enzymes.
Question
BXS4NIE8Nyqr/vr2cqpfIxnOi8TZODuqIE9JHK7fBYWlA5YZB8B8p67d1k2S+jkZwTjL+yXRlG4mqslYL/CP+R7bPfHnNx0RPAiesxe27vJubq5hRzVv8prpzsw/d41TOQqxIfkPL4FnjAo53m/Q97T3vAyfvAOCsgle6yg5l/IlzU9+HJ/5DoxiyWLcj++8IEwUkMeGvWwwIWjPU/tD40FdwOhMrfaBxt/qe35J6DkF6YL19DocYwoURU+aOsKDiJx0oHowPfmoKZjYlW/yF+lIpxC5lokWeRDYdower4dCJxJUdBJ1zTKrnLWPeDZ/X9Fi4F0iNWe0oegjaAT8wYyhiqSXCWmT3NT8aAAWuu02XbQLExz2rNwFrUZ5ApQPgR6V/worQ0Rn5srHCmlQlrbrkbjVEn+4ffgkFZ397KEPYZ5YYbEJhP6kwVEuMs6Ne07YhACuYhqpBfILHj2I3N/oVc1OXplrdHXFrOd3MztdSVURQqsmIUWncNWe09Sbnl2tIBAdCnJAETm1+VbD4XkOMnLNU5fyUmTqZ1hmgtIo1KHC9jQZEj07P8q/gJDR0GnxtsIRUQ26yeNZWRXj8BoxTlNOJZI8fFZHjJXv8rFaqa/cViuLcdLW8+JeyG0HkOAdQ2hRCFyEOza1dZN2jdMZSZJMD9yWsct5MW22C49wV85kPSoznJywh9r6+ix6XDjKB3Ny0hwdzpqsLHnRHjmKQ5fKr8txvyj3cBzJ/0mNFO71EtXrnZWiXGQh3QvZ5yHxsDBH9pyVsGQhYO9bSY0HdaJeJEJXrOK7WsppZYwO+qQn3mAnTvtC8yKbbGbELoA92ggsOKhIakKpMqn1Enb3mY7is9bIwbDqFAL7hm2sRj9Qy9s7o1+2LuzX/PheVo17qcCHfWA5h6/JBYdOnmQd5Fl0ly/f8EFQfIWUx6MsW/kBxHB+RPudhwgR/o7PSglnfRMjT4sYaZeawsL4iSa6e1sZCSHW0MCrtLx05TrzdLPR2VwZuuJM5ewjb912hk7Ik4OJOrVsfVmPb+8S2wpYAPZALGvXBguxdA1Z7wxVPEyCKrQU94C7MrJ75BHtNUmqt8Ull2t+bDaZ/z6z1ASV+zubdgtwE2PS7/7/mmr95PkM/M3lcitrnm7v6JYDrjVkNx0kUOSam/n4T+yuPQtGjrCAM536xWj2p5cTIAVhhY7tfHeWBPmyT9Is922U4w1IavtjjauIBUMJWUS77Alc4yQXn23zkNdId9ouEuYHq+0j8+jYtYoCSH/gGFW8Instant TA
Which fragment size appears in the “EcoRI only” digest and not the double digest?
Question
Gp0R8GGG51AzGgwZgJBJq6lj3eTknjzP2Cr2KitQeJd+RZUDwGC3tAWRGlM/5UX901eXA3lOYrs6ZVWbVk4FQqb93msWvmeZe0Qi+6yT+0UicUp+10HB0NqWcF/l20MLqEeQvVluVy0ABwXBK/ceoSZVVflVk5lxRtvh83OhKmqpt7YjQSSK03JQLN585dhn1kSucTuaJf/D4CbP3p0t7/HiHuAg2PUt+EbsavIeNVCL2gDn8pSfKc050EcNGyrAqZo1WsU7T/r6Mzr0kMriRyyyhQJv02QqxMmL51R4oknbl6N6Rxn9kupfIk2FwPiQ0oLVbyF1JTfimMe3oTvUN+RMy1/Jr7zWy7djugyUERpiUkXwgITwGwFpcKl2CJmYfR9jOk+YDbvdRDsVtEk0qQ1Rk3m0dUOJBgYXjFgM8MUZkk9E8XRe5+JesBBOiyyiWkTKOp02HbbuRbdFPCe4QjYHLppAS1o/I9iHC1rKJApua8q6JI7Jv6X5aVpkXRVtg7uVlUN8gtbtt8lnPpeXOIhjka55v2XZPIrzk54Ff5HQq0KXzbBDKv0YklMO3v5m87EIDHy4QC2FxWe3MTm64Md7Qk0BsJVmDtnmm6CrFTHd8snoMkjHHggXAAnaSiAgtDeoJtNQ0lo=Instant TA
Since you are looking for the EcoRI fragment that would be cut, some combination of the double digest fragment sizes should total the size of one of the Eco fragments from the single digest.
Question
mihouX2tRerbti1OB2EZykL9RFdCX1VTlC/CENOR+vh6BHHvIWqVcdfCHS8Uozj8jPBQEpHr67vCoPRsMQoLT4pQPTDtFB/68KLyV7K58tTXMlzruV8zH7oXXB5lxm6V4v6U71cRRW7BecNrLoHAeXCqodC55ttFJYAaXUJwAMZr65TsE61dfEMZfHbHdHW8jn24/DHv7JQQbMkwTQPISnXFjaYe4rjKYqfsq7sQ8yOP5ELea1R72CA76d7hYNkKji1sK1AaHAkon4CU/kDPLaCRdqzqIX1GBVHJB1dKihPkD5cbED0ETZnzaxZUfwkV4MK3HbPv2NMPF+Q7t5G54aPetNiTQofO8T4kditpIXy3NiH6BH3m+vvEMf43QFtu1gTSDS/WUSdHsNPRH1U3YHqkb4HHfT5UwNxCoWSnXjebHIi8chQG2JRQcv4mXDrYsmM26Xwok7fF7t6dcM2d2evWF7IJ/4xRCjTOfHnb9QuPVEidXrwGX0a3m+2IzIjNkgvKJGb9+QH6JvU5Instant TA
These two fragments are additive when combined. They would add to the size of the EcoRI fragment.
1.6 Step 4: Solving the Problem
Question
1. Draw a restriction map based on the third lane of the gel. Choose from the dropdown menus below to correctly label each restriction enzyme recognition site and fragment size.

| A. jAW2nq2QxNaPmtooReC4oNUUyKG/fW3ybjmW74MKvUE6jD84WLaNITJx8ki/7SZi1Gvca4iV4dxkOG38GKeoToo58tR4RVN433Ns/it5yWo= | B. 4IFBf9ScuaDHv4T88pKVLY5cyOOOx0XKqZbHdQHyd2kPNeFTaACHltwZ7ojhpdwQUSPOWWdaiHx9rq6xLJEZUqToMduwogkP9Dmsd57wJTk= | C. TN8pRmL1WdrhZ2x57C7AmhEk+u50Sg/Tak0OuZzK+DWIJwaICEahWdviPD9p5FPXXwe+KeU3yja6aqx0q9P7VsSjnn3l1cLmdDhRf9Afwk0= | D. p8CSlrXINuswoOcdDxwvC9HTT5XW6BVO047taLVdelAuzYN49m3UReW0PPI5CpB+xLnsTw8O8x8Kyqxofd3v69FTcOdbs7nIMSzpCSnEKRI= | E. pWZZxi3nhJchInvjqYiiq2rCwh/dPk0r6zgH4BX2Fh7YSmnpXPJmXxP3SXF+qdyszYFd31aCw2SFY5m6PZ2kYeDgLAwVszy1bgJC3bOdD0s= | F. 4IFBf9ScuaDHv4T88pKVLY5cyOOOx0XKqZbHdQHyd2kPNeFTaACHltwZ7ojhpdwQUSPOWWdaiHx9rq6xLJEZUqToMduwogkP9Dmsd57wJTk= | G. bxslCdwuwP4Bj31zlKqGeC4guY+f6ohY42FeTaqmETT3oTntx+8DTcWd+pgGrs2LDIZcpgQbYja//ttI42YlKj3BlVHPl9f1Zc80Mr82YUA= |
Instant TA
Remember that there are EcoRI fragments which are not cut by BamHI. These fragment sizes should not contain a BamHI restriction site on your map. Begin your mapping by identifying the largest EcoRI generated fragment—remember, this fragment contains a BamHI site. Be sure that you are locating the 7 kb and 1 kb fragments generated by a BamHI enzyme next to each other on the map. Keep in mind that the total size of the fragments needs to be equal to the size of the initial DNA fragment before digestion.