12.4 Eukaryotic DNA Replication Is Similar to Bacterial Replication but Differs in Several Aspects
Although eukaryotic replication resembles bacterial replication in many respects, replication in eukaryotic cells presents several additional challenges. First, the much greater size of eukaryotic genomes requires that replication be initiated at multiple origins. Second, eukaryotic chromosomes are linear, whereas prokaryotic chromosomes are circular. Third, the DNA template is associated with histone proteins in the form of nucleosomes, and nucleosome assembly must immediately follow DNA replication.
Eukaryotic Origins
Researchers first isolated eukaryotic origins of replication from yeast cells by demonstrating that certain DNA sequences confer the ability to replicate when transferred from a yeast chromosome to small circular pieces of DNA (plasmids). These autonomously replicating sequences (ARSs) enabled any DNA to which they were attached to replicate. They were subsequently shown to be the origins of replication in yeast chromosomes. The origins of replication of different eukaryotic organisms vary greatly in sequence, although they usually contain a number of A-T base pairs. A multiprotein complex, the origin-recognition complex (ORC), binds to origins and unwinds the DNA in this region.
CONCEPTS
Eukaryotic DNA contains many origins of replication. At each origin, a multiprotein origin-recognition complex binds to initiate the unwinding of the DNA.
 CONCEPT CHECK 8
CONCEPT CHECK 8
In comparison with prokaryotes, what are some differences in the genome structure of eukaryotic cells that affect how replication takes place?
341
The Licensing of DNA Replication
Eukaryotic cells utilize thousands of origins, and so the entire genome can be replicated in a timely manner. The use of multiple origins, however, creates a special problem in the timing of replication: the entire genome must be precisely replicated once and only once in each cell cycle so that no genes are left unreplicated and no genes are replicated more than once. How does a cell ensure that replication is initiated at thousands of origins only once per cell cycle?
The precise replication of DNA is accomplished by the separation of the initiation of replication into two distinct steps. In the first step, the origins are licensed—approved for replication. This step takes place early in the cell cycle when a replication licensing factor attaches to an origin. In the second step, the replication machinery initiates replication at each licensed origin. The key is that the replication machinery functions only at licensed origins. As the replication forks move away from the origin, the licensing factor is removed, leaving the origin in an unlicensed state, where replication cannot be initiated again until the license is renewed. To ensure that replication takes place only once per cell cycle, the licensing factor is active only after the cell has completed mitosis and before the replication is initiated.
One eukaryotic licensing factor is a complex called MCM (for minichromosome maintenance), which contains a DNA helicase that unwinds a short stretch of DNA in the initiation of replication. MCM must bind to the DNA for replication to initiate at an origin. After replication has begun at an origin, a protein called Geminin prevents MCM from binding to DNA and reinitiating replication at that origin. At the end of mitosis, Geminin is degraded, allowing MCM to bind once again to DNA and relicense the origin. MCM also functions as the DNA helicase during the replication process.
Unwinding
Several different helicases that separate double-stranded DNA have been isolated from eukaryotic cells, as have single-strand-binding proteins and topoisomerases (which have a function equivalent to the DNA gyrase in bacterial cells). These enzymes and proteins are assumed to function in unwinding eukaryotic DNA in much the same way as their bacterial counterparts do.
Eukaryotic DNA Polymerases
Some significant differences in the processes of bacterial and eukaryotic replication are in the number and functions of DNA polymerases. Eukaryotic cells contain a number of different DNA polymerases that function in replication, recombination, and DNA repair.
Three DNA polymerases carry out most of nuclear DNA synthesis during replication: DNA polymerase α, DNA polymerase δ, and DNA polymerase ε (Table 12.5).
| DNA Polymerase | 5′→3′ Polymerase Activity | 3′→5′ Exonuclease Activity | Cellular Function |
|---|---|---|---|
| α (alpha) | Yes | No | Initiation of nuclear DNA synthesis and DNA repair; has primase activity |
| δ (delta) | Yes | Yes | Lagging-strand synthesis of nuclear DNA, DNA repair, and translesion DNA synthesis |
| ε (epsilon) | Yes | Yes | Leading-strand synthesis |
| γ (gamma) | Yes | Yes | Replication and repair of mitochondrial DNA |
| ξ (zeta) | Yes | No | Translesion DNA synthesis |
| η (eta) | Yes | No | Translesion DNA synthesis |
| θ (theta) | Yes | No | DNA repair |
| ι (iota) | Yes | No | Translesion DNA synthesis |
| κ (kappa) | Yes | No | Translesion DNA synthesis |
| λ (lambda) | Yes | No | DNA repair |
| μ (mu) | Yes | No | DNA repair |
| σ (sigma) | Yes | No | Nuclear DNA replication (possibly), DNA repair, and sister-chromatid cohesion |
| φ (phi) | Yes | No | Translesion DNA synthesis |
| Rev1 | Yes | No | DNA repair |
| Note: The polymerases listed at the top of the table are those that carry out DNA replication. | |||
342
DNA polymerase α contains primase activity and initiates nuclear DNA synthesis by synthesizing an RNA primer, followed by a short string of DNA nucleotides. After DNA polymerase α has laid down from 30 to 40 nucleotides, DNA polymerase δ completes replication on the lagging strand. Similar in structure and function to DNA polymerase δ, DNA polymerase ε replicates the leading strand. Other DNA polymerases take part in repair and recombination or catalyze the replication of organelle DNA.
Some DNA polymerases, such as DNA polymerase δ and DNA polymerase ε, are capable of replicating DNA at high speed and with high fidelity (few mistakes) because they have active sites that snugly and exclusively accommodate the four normal DNA nucleotides, adenosine, guanosine, cytidine, and thymidine monophosphates. As a result of this specificity, distorted DNA templates and abnormal bases are not readily accommodated within the active site of the enzyme. When these errors are encountered in the DNA template, the high-fidelity DNA polymerases stall and are unable to bypass the lesion.
Other DNA polymerases have lower fidelity but are able to bypass distortions in the DNA template. These specialized translesion DNA polymerases generally have a more open active site and are able to accommodate and copy templates with abnormal bases, distorted structures, and bulky lesions. Thus, these specialized enzymes can bypass such errors but, because their active sites are more open and accommodating, they tend to make more errors. In replication, high-speed, high-fidelity enzymes are generally used until they encounter a replication block. At that point, one or more of the translesion polymerases takes over, bypasses the lesion, and continues replicating a short section of DNA. Then, the translesion polymerases detach from the replication fork and high-fidelity enzymes resume replication with high speed and accuracy. DNA-repair enzymes often repair errors produced by the translesion polymerases, although some of these errors may escape detection and lead to mutations.
CONCEPTS
There are a large number of different DNA polymerases in eukaryotic cells. DNA polymerases α, δ, and ε carry out replication on the leading and lagging strands. Other DNA polymerases carry out DNA repair. Specialized translesion polymerases are used to bypass distortions of the DNA template that normally stall the main DNA polymerases.
 CONCEPT CHECK 9
CONCEPT CHECK 9
Some of the eukaryotic DNA polymerases have a tendency to make errors in replication. Why would a cell use an error-prone DNA polymerase instead of one that is more accurate?
Nucleosome Assembly
Eukaryotic DNA is complexed to histone proteins in nucleosome structures that contribute to the stability and packing of the DNA molecule (see Figure 11.4). In replication, chromatin structure is disrupted by the replication fork, but nucleosomes are quickly reassembled on the two new DNA molecules. Electron micrographs of eukaryotic DNA, such as that in Figure 12.16, show recently replicated DNA already covered with nucleosomes, indicating that nucleosomes are reassembled quickly.

The creation of new nucleosomes requires three steps: (1) the disruption of the original nucleosomes on the parental DNA molecule ahead of the replication fork; (2) the redistribution of preexisting histones on the new DNA molecules; and (3) the addition of newly synthesized histones to complete the formation of new nucleosomes. Before replication, a single DNA molecule is associated with histone proteins. After replication and nucleosome assembly, two DNA molecules are associated with histone proteins. Do the original histones of a nucleosome remain together, attached to one of the new DNA molecules, or do they disassemble and mix with new histones on both DNA molecules?
343
Techniques similar to those employed by Meselson and Stahl to determine the mode of DNA replication were used to address this question. Cells were cultivated for several generations in a medium containing amino acids labeled with a heavy isotope. The histone proteins incorporated these heavy amino acids and were dense (Figure 12.17). The cells were then transferred to a culture medium that contained amino acids labeled with a light isotope. Histones assembled after the transfer possessed the new, light amino acids and were less dense.
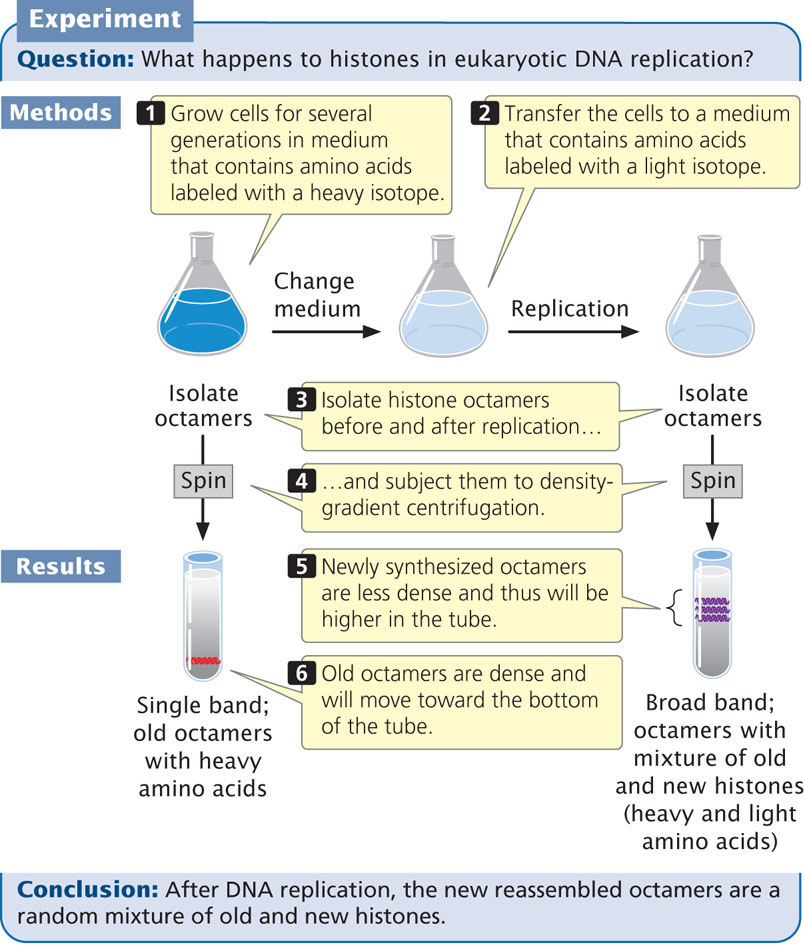
After replication, the histone octamers were isolated and centrifuged in a density gradient. Results showed that, after replication, the octamers were in a continuous band between high density (representing old octamers) and low density (representing new octamers). This finding indicates that newly assembled octamers consist of a mixture of old and new histones. Further evidence indicates that reconstituted nucleosomes appear on the new DNA molecules quickly after the new DNA emerges from the replication machinery.
The reassembly of nucleosomes during replication is facilitated by proteins called histone chaperones, which are associated with the helicase enzyme that unwinds the DNA. The histone chaperones accept old histones from the original DNA molecule and deposit them, along with newly synthesized histones, on the two new DNA molecules. Current evidence suggests that the original nucleosome is broken down into two H2A-H2B dimers (each dimer consisting of one H2A and one H2B) and a single H3-H4 tetramer (each tetramer consisting of two H3 histones and two H4 histones). The old H3-H4 tetramer is then transferred randomly to one of the new DNA molecules and serves as a foundation onto which either new or old copies of H2A-H2B dimers are added. Newly synthesized H3-H4 tetramers and H2A-H2b dimers also are added to each new DNA molecule to complete the formation of new nucleosomes. The assembly of the new nucleosomes is facilitated by a protein called chromatin-assembly factor 1 (CAF-1).  TRY PROBLEM 23
TRY PROBLEM 23
CONCEPTS
After DNA replication, new nucleosomes quickly reassemble on the molecules of DNA. Nucleosomes break down in the course of replication and reassemble from a mixture of old and new histones. The reassembly of nucleosomes during replication is facilitated by histone chaperones and chromatin-assembly factors.
The Location of Replication Within the Nucleus
The DNA polymerases that carry out replication are frequently depicted as moving down the DNA template, much as a locomotive travels along a train track. Recent evidence suggests that this view is incorrect. A more accurate view is that the polymerase is fixed in location and template DNA is threaded through it, with newly synthesized DNA molecules emerging from the other end.
Techniques of fluorescence microscopy, which are able to reveal active sites of DNA synthesis, show that most replication in the nucleus of a eukaryotic cell takes place at a limited number of fixed sites, often referred to as replication factories. Time-lapse micrographs reveal that newly duplicated DNA is extruded from these particular sites. Similar results have been obtained for bacterial cells.
DNA Synthesis and the Cell Cycle
In rapidly dividing bacteria, DNA replication is continuous. In eukaryotic cells, however, replication is coordinated with the cell cycle. Passage through the cell cycle, including the onset of replication, is controlled by cell-cycle checkpoints. The important G1/S checkpoint (see Chapter 2) holds the cell cycle in G1 until the DNA is ready to be replicated. After the G1/S checkpoint is passed, the cell enters S phase and the DNA is replicated. The replication licensing system then ensures that the DNA is not replicated again until after the cell has passed through mitosis.
344
Replication at the Ends of Chromosomes
A fundamental difference between eukaryotic and bacterial replication arises because eukaryotic chromosomes are linear and thus have ends. As already stated, the 3′-OH group needed for replication by DNA polymerases is provided at the initiation of replication by RNA primers that are synthesized by primase. This solution is temporary because, eventually, the primers must be removed and replaced by DNA nucleotides. In a circular DNA molecule, elongation around the circle eventually provides a 3′-OH group immediately in front of the primer (Figure 12.18a). After the primer has been removed, the replacement DNA nucleotides can be added to this 3′-OH group.
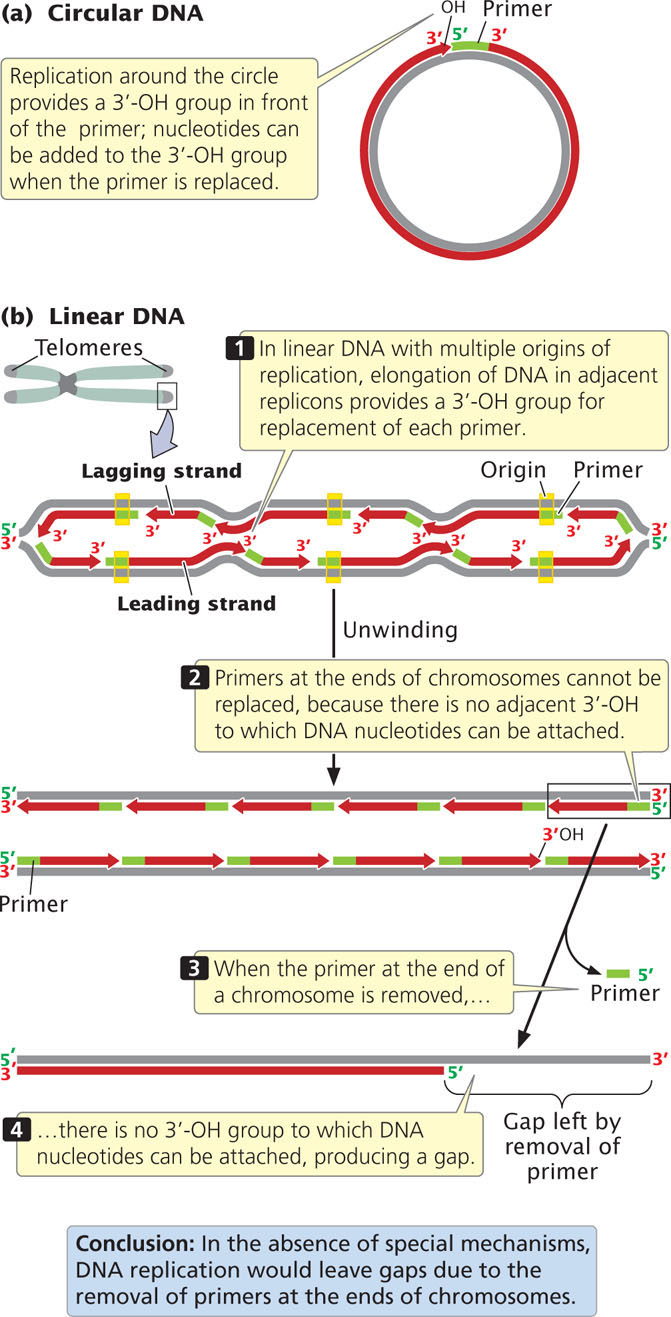
The End-Replication Problem
In linear chromosomes with multiple origins, the elongation of DNA in adjacent replicons also provides a 3′-OH group preceding each primer (Figure 12.18b). At the very end of a linear chromosome, however, there is no adjacent stretch of replicated DNA to provide this crucial 3′-OH group. When the primer at the end of the chromosome has been removed, it cannot be replaced by DNA nucleotides, which produces a gap at the end of the chromosome, suggesting that the chromosome should become progressively shorter with each round of replication. Chromosome shortening would mean that, when an organism reproduced, it would pass on shorter chromosomes than it had inherited. Chromosomes would become shorter with each new generation and would eventually destabilize. This situation has been termed the end-replication problem. Chromosome shortening does in fact take place in many somatic cells but, in single-celled organisms, germ cells, and early embryonic cells, chromosomes do not shorten and self-destruct. So how are the ends of linear chromosomes replicated?
Telomeres and Telomerase
The ends of chromosomes—the telomeres—possess several unique features, one of which is the presence of many copies of a short repeated sequence. In the protozoan Tetrahymena (where these repeated sequences were first discovered) this telomeric repeat is TTGGGG (see Table 11.2), with this G-rich strand typically protruding beyond the C-rich strand (Figure 12.19a; also see the section on Telomere Structure in Chapter 11):
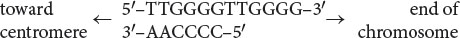
The single-stranded protruding end of the telomere, known as the G overhang can be extended by telomerase, an enzyme with both a protein and an RNA component (also known as a ribonucleoprotein). The RNA part of the enzyme contains from 15 to 22 nucleotides that are complementary to the sequence on the G-rich strand. This sequence pairs with the overhanging 3′ end of the DNA (Figure 12.19b) and provides a template for the synthesis of additional DNA copies of the repeats. DNA nucleotides are added to the 3′ end of the strand one at a time (Figure 12.19c) and, after several nucleotides have been added, the RNA template moves down the DNA and more nucleotides are added to the 3′ end (Figure 12.19d). Usually, from 14 to 16 nucleotides are added to the 3′ end of the G-rich strand.
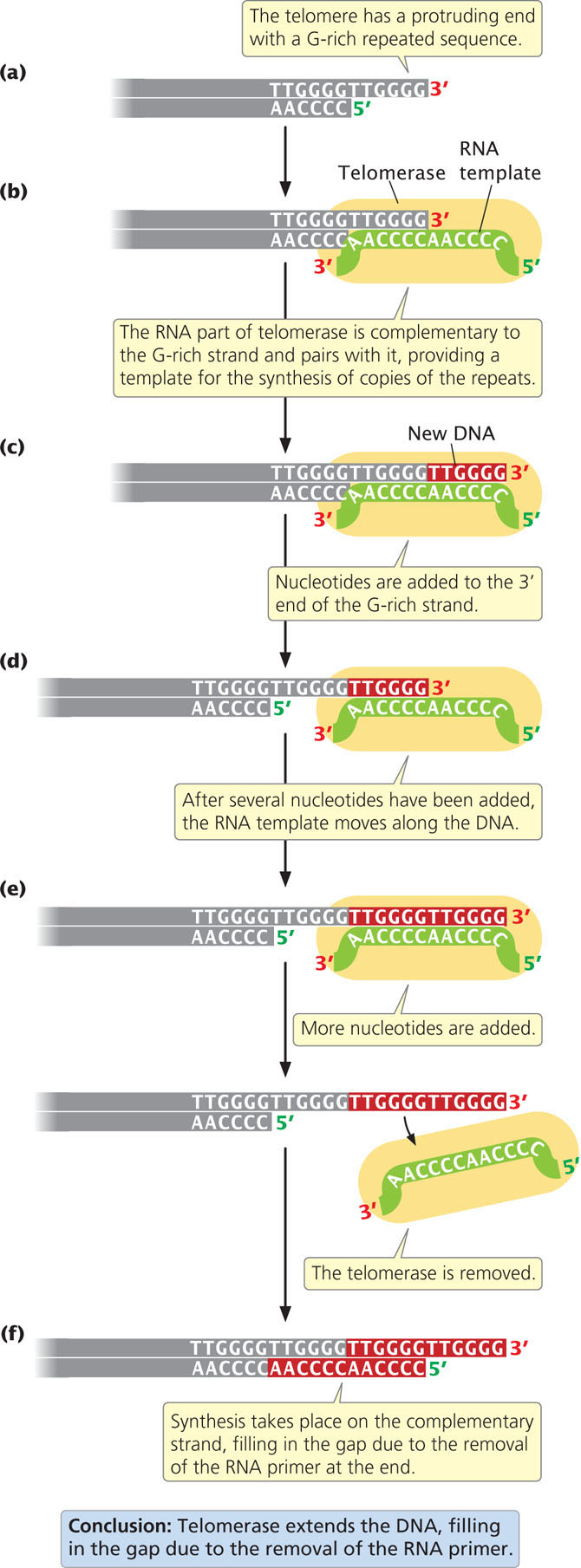
In this way, the telomerase can extend the 3′ end of the chromosome without the use of a complementary DNA template (Figure 12.19e). How the complementary C-rich strand is synthesized (Figure 12.19f) is not clear. It may be synthesized by conventional replication, with DNA polymerase α synthesizing an RNA primer on the 5′ end of the extended (G-rich) template. The removal of this primer once again leaves a gap at the 5′ end of the chromosome, but this gap does not matter, because the end of the chromosome is extended at each replication by telomerase; so, the chromosome does not become shorter overall.
345
Telomerase is present in single-celled organisms, germ cells, early embryonic cells, and certain proliferative somatic cells (such as bone-marrow cells and cells lining the intestine), all of which must undergo continuous cell division. Most somatic cells have little or no telomerase activity, and chromosomes in these cells progressively shorten with each cell division. These cells are capable of only a limited number of divisions; when the telomeres have shortened beyond a critical point a chromosome becomes unstable, has a tendency to undergo rearrangements, and is degraded. These events lead to cell death.
CONCEPTS
The ends of eukaryotic chromosomes are replicated by an RNA-protein enzyme called telomerase. This enzyme adds extra nucleotides to the G-rich DNA strand of the telomere.
 CONCEPT CHECK 10
CONCEPT CHECK 10
What would be the result if an organism’s telomerase were mutated and nonfunctional?
- No DNA replication would take place.
- The DNA polymerase enzyme would stall at the telomere.
- Chromosomes would shorten with each new generation.
- RNA primers could not be removed.
Telomerase, Aging, and Disease
The shortening of telomeres may contribute to the process of aging. The telomeres of genetically engineered mice that lack a functional telomerase gene (and therefore do not express telomerase in somatic or germ cells) undergo progressive shortening in successive generations. After several generations, these mice show some signs of premature aging, such as graying, hair loss, and delayed wound healing. Through genetic engineering, it is also possible to create somatic cells that express telomerase. In these cells, telomeres do not shorten, cell aging is inhibited, and the cells will divide indefinitely.
Some of the strongest evidence that telomere length is related to aging comes from studies of telomeres in birds. In 2012, scientists in the United Kingdom measured telomere length in red blood cells taken from 99 zebra finches at various times during their lives. The scientists found a strong correlation between telomere length and longevity: birds with longer telomeres lived longer than birds with short telomeres. The strongest predictor of life span was when telomere length was measured early in life, at 25 days, which is roughly equivalent to human adolescence. Although these observations suggest that telomere length is associated with aging in some animals, the precise role of telomeres in human aging remains uncertain.
346
Some diseases are associated with abnormalities of telomere replication. People who have Werner syndrome, an autosomal recessive disease, show signs of premature aging that begins in adolescence or early adulthood, including wrinkled skin, graying of the hair, baldness, cataracts, and muscle atrophy. They often develop cancer, osteoporosis, heart and artery disease, and other ailments typically associated with aging. The causative gene, WRN, has been mapped to human chromosome 8 and normally encodes a RecQ helicase enzyme. This enzyme is necessary for the efficient replication of telomeres. In people who have Werner syndrome, this helicase is defective and, consequently, the telomeres shorten prematurely.
Another disease associated with abnormal maintenance of telomeres is dyskeratosis congenita, which leads to progressive bone-marrow failure, in which the bone marrow fails to produce enough new blood cells. People with an X-linked form of the disease have a mutation in a gene that encodes dyskerin, a protein that normally helps process the RNA component of telomerase. People who have the disease typically inherit short telomeres from a parent who carries the mutation and who is unable to maintain telomere length in his or her germ cells owing to defective dyskerin. In families that carry this mutation, telomere length typically shortens with each successive generation, leading to anticipation, a progressive increase in the severity of the disease over generations (see Chapter 5).
Telomerase also appears to play a role in cancer. Cancer tumor cells have the capacity to divide indefinitely, and the telomerase enzyme is expressed in 90% of all cancers. Some recent evidence indicates that telomerase may stimulate cell proliferation independently of its effect on telomere length, and so the mechanism by which telomerase contributes to cancer is not clear. As will be discussed in Chapter 23, cancer is a complex, multistep process that usually requires mutations in at least several genes. Telomerase activation alone does not lead to cancerous growth in most cells, but it does appear to be required, along with other mutations, for cancer to develop. Some experimental cancer drugs work by inhibiting the action of telomerase.
One of the difficulties in studying the effect of telomere shortening on the aging process is that the expression of telomerase in somatic cells also promotes cancer, which may shorten a person’s life span. To circumvent this problem, Antonia Tomas-Loba and her colleagues created genetically engineered mice that expresssed telomerase and carried genes that made them resistant to cancer. These mice had longer telomeres, lived longer, and exhibited fewer age-related changes, such as skin alterations, a decrease in neuromuscular coordination, and degenerative diseases. These results support the idea that telomere shortening contributes to aging.  TRY PROBLEM 35
TRY PROBLEM 35
Replication in Archaea
The process of replication in archaea has a number of features in common with replication in eukaryotic cells; many of the proteins taking part are more similar to those in eukaryotic cells than to those in eubacteria. Like eubacteria, some archaea have a single replication origin, but the archaean Sulfolobus solfataricus has two origins of replication, similar to the multiple origins seen in eukaryotic genomes. The replication origins of archaea do not contain the typical sequences recognized by bacterial initiator proteins; instead, they have sequences that are similar to those found in eukaryotic origins. The initiator proteins of archaea also are more similar to those of eukaryotes than to those of eubacteria. These similarities in replication between archaeal and eukaryotic cells reinforce the conclusion that the archaea are more closely related to eukaryotic cells than to the prokaryotic eubacteria.