4.6Amino Acids Are Encoded by Groups of Three Bases Starting from a Fixed Point
Amino Acids Are Encoded by Groups of Three Bases Starting from a Fixed Point
The genetic code is the relation between the sequence of bases in DNA (or its RNA transcripts) and the sequence of amino acids in proteins. Experiments by Marshall Nirenberg, Har Gobind Khorana, Francis Crick, Sydney Brenner, and others established the following features of the genetic code by 1961:
1. Three nucleotides encode an amino acid. Proteins are built from a basic set of 20 amino acids, but there are only four bases. Simple calculations show that a minimum of three bases is required to encode at least 20 amino acids. Genetic experiments showed that an amino acid is in fact encoded by a group of three bases, or codon.
2. The code is nonoverlapping. Consider a base sequence ABCDEF. In an overlapping code, ABC specifies the first amino acid, BCD the next, CDE the next, and so on. In a nonoverlapping code, ABC designates the first amino acid, DEF the second, and so forth. Genetic experiments again established the code to be nonoverlapping.
3. The code has no punctuation. In principle, one base (denoted as Q) might serve as a “comma” between groups of three bases.
… QABCQDEFQGHIQJKLQ …
However, it is not the case. Rather, the sequence of bases is read sequentially from a fixed starting point, without punctuation.
4. The code has directionality. The code is read from the 5′ end of the messenger RNA to its 3′ end.
125
5. The genetic code is degenerate. Most amino acids are encoded by more than one codon. There are 64 possible base triplets and only 20 amino acids, and in fact 61 of the 64 possible triplets specify particular amino acids. Three triplets (called stop codons) designate the termination of translation. Thus, for most amino acids, there is more than one code word.
Major features of the genetic code
All 64 codons have been deciphered (Table 4.5). Because the code is highly degenerate, only tryptophan and methionine are encoded by just one triplet each. Each of the other 18 amino acids is encoded by two or more. Indeed, leucine, arginine, and serine are specified by six codons each.
|
First Position (5′ end) |
Second Position |
Third Position (3′ end) |
|||
|---|---|---|---|---|---|
|
U |
C |
A |
G |
||
|
U |
Phe |
Ser |
Tyr |
Cys |
U |
|
Phe |
Ser |
Tyr |
Cys |
C |
|
|
Leu |
Ser |
Stop |
Stop |
A |
|
|
Leu |
Ser |
Stop |
Trp |
G |
|
|
C |
Leu |
Pro |
His |
Arg |
U |
|
Leu |
Pro |
His |
Arg |
C |
|
|
Leu |
Pro |
Gln |
Arg |
A |
|
|
Leu |
Pro |
Gln |
Arg |
G |
|
|
A |
Ile |
Thr |
Asn |
Ser |
U |
|
Ile |
Thr |
Asn |
Ser |
C |
|
|
Ile |
Thr |
Lys |
Arg |
A |
|
|
Met |
Thr |
Lys |
Arg |
G |
|
|
G |
Val |
Ala |
Asp |
Gly |
U |
|
Val |
Ala |
Asp |
Gly |
C |
|
|
Val |
Ala |
Glu |
Gly |
A |
|
|
Val |
Ala |
Glu |
Gly |
G |
|
|
Note: This table identifies the amino acid encoded by each triplet. For example, the codon 5′- AUG- |
|||||
Codons that specify the same amino acid are called synonyms. For example, CAU and CAC are synonyms for histidine. Note that synonyms are not distributed haphazardly throughout the genetic code. In Table 4.5, an amino acid specified by two or more synonyms occupies a single box (unless it is specified by more than four synonyms). The amino acids in a box are specified by codons that have the same first two bases but differ in the third base, as exemplified by GUU, GUC, GUA, and GUG. Thus, most synonyms differ only in the last base of the triplet. Inspection of the code shows that XYC and XYU always encode the same amino acid, and XYG and XYA usually encode the same amino acid as well. The structural basis for these equivalences of codons becomes evident when we consider the nature of the anticodons of tRNA molecules (Section 30.3).
What is the biological significance of the extensive degeneracy of the genetic code? If the code were not degenerate, 20 codons would designate amino acids and 44 would lead to chain termination. The probability of mutating to chain termination would therefore be much higher with a non-
126
Messenger RNA contains start and stop signals for protein synthesis
Messenger RNA is translated into proteins on ribosomes—large molecular complexes assembled from proteins and ribosomal RNA. How is mRNA interpreted by the translation apparatus? The start signal for protein synthesis is complex in bacteria. Polypeptide chains in bacteria start with a modified amino acid—
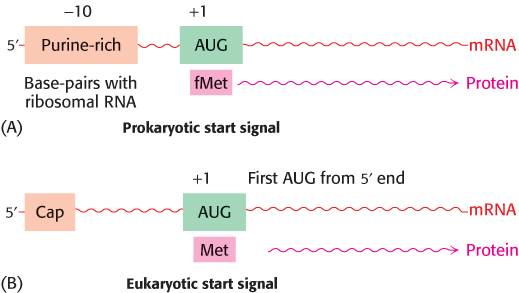
As already mentioned, UAA, UAG, and UGA designate chain termination. These codons are read not by tRNA molecules but rather by specific proteins called release factors (Section 30.3). Binding of a release factor to the ribosome releases the newly synthesized protein.
The genetic code is nearly universal
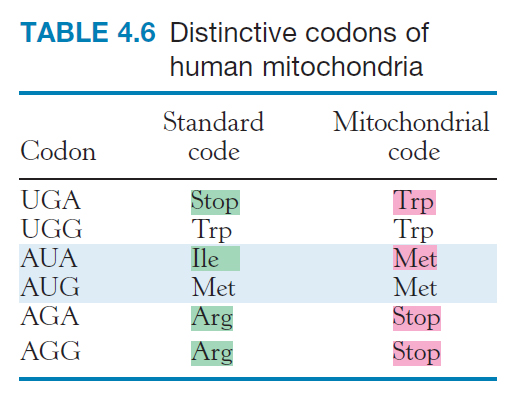
Most organisms use the same genetic code. This universality accounts for the fact that human proteins, such as insulin, can be synthesized in the bacterium E. coli and harvested from it for the treatment of diabetes. However, genome-
Why has the code remained nearly invariant through billions of years of evolution, from bacteria to human beings? A mutation that altered the reading of mRNA would change the amino acid sequence of most, if not all, proteins synthesized by that particular organism. Many of these changes would undoubtedly be deleterious, and so there would be strong selection against a mutation with such pervasive consequences.
127