35.4A Rotary Motor Drives Bacterial Motion
A Rotary Motor Drives Bacterial Motion
In 1 s, a motile bacterium can move approximately 25 μm, or about 10 body lengths. A human being sprinting at a proportional rate would complete the 100-
Bacteria swim by rotating their flagella
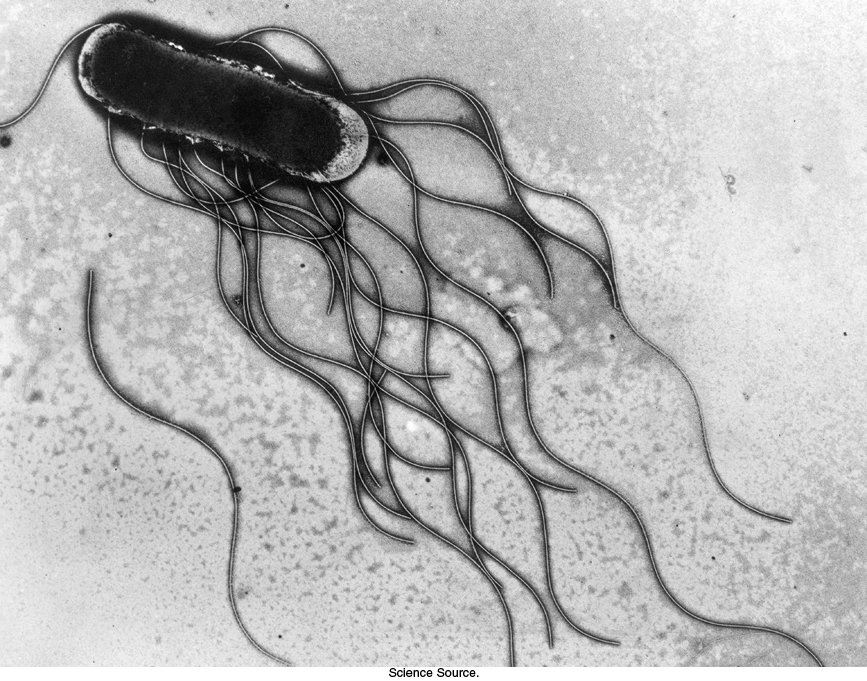
Bacteria such as Escherichia coli and Salmonella typhimurium swim by rotating flagella that lie on their surfaces (Figure 35.25). When the flagella rotate in a counterclockwise direction (viewed from outside a bacterium), the separate flagella form a bundle that very efficiently propels the bacterium through solution.
Bacterial flagella are polymers approximately 15 nm in diameter and as much as 15 μm in length, composed of 53-
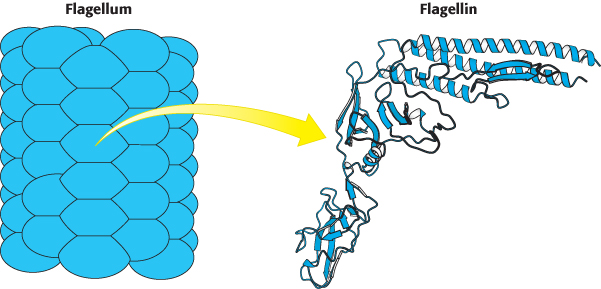
 FIGURE 35.26 Structure of flagellin. A bacterial flagellum is a helical polymer of the protein flagellin. Notice that each subunit corresponds to a bent structure with a relatively flat surface facing the hollow core of the flagellum.
FIGURE 35.26 Structure of flagellin. A bacterial flagellum is a helical polymer of the protein flagellin. Notice that each subunit corresponds to a bent structure with a relatively flat surface facing the hollow core of the flagellum.
Proton flow drives bacterial flagellar rotation
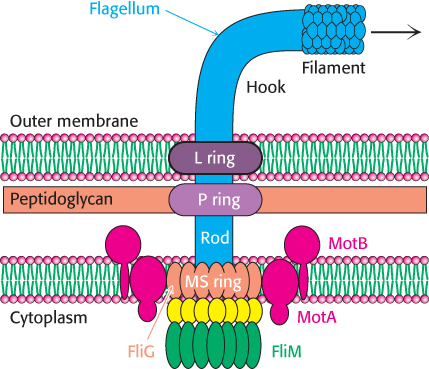
Early experiments by Julius Adler demonstrated that ATP is not required for flagellar motion. What powers these rotary motors? The necessary free energy is derived from the proton gradient that exists across the plasma membrane. The flagellar motor is quite complex, containing as many as 40 distinct proteins (Figure 35.27). Five components particularly crucial to motor function have been identified through genetic studies. MotA is a membrane protein that appears to have four transmembrane helices as well as a cytoplasmic domain. MotB is another membrane protein with a single trans-
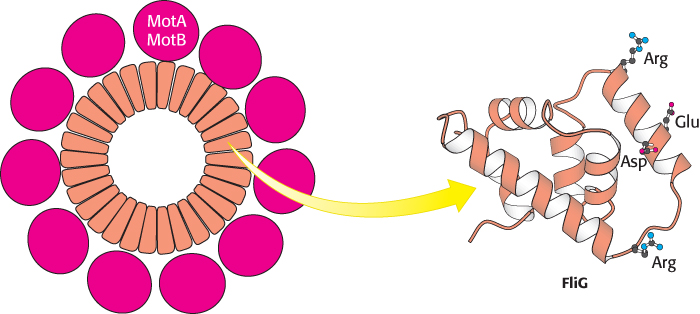
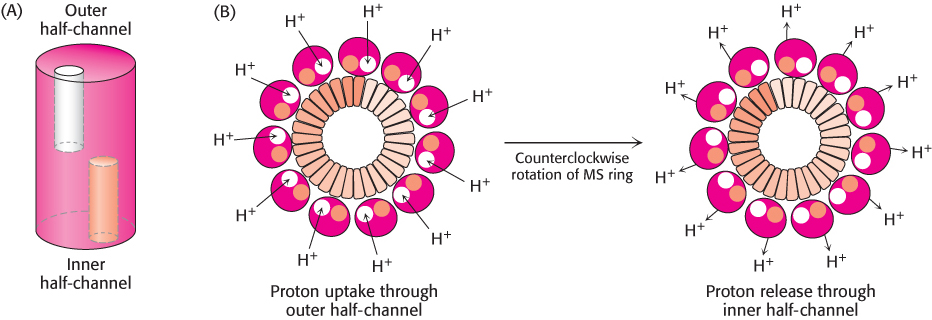
 FIGURE 35.28 Flagellar motor components. Approximately 30 subunits of FliG assemble to form part of the MS ring. The ring is surrounded by approximately 11 structures consisting of MotA and MotB. Notice that the carboxyl-
FIGURE 35.28 Flagellar motor components. Approximately 30 subunits of FliG assemble to form part of the MS ring. The ring is surrounded by approximately 11 structures consisting of MotA and MotB. Notice that the carboxyl-1027
The MotA–
1028
Bacterial chemotaxis depends on reversal of the direction of flagellar rotation
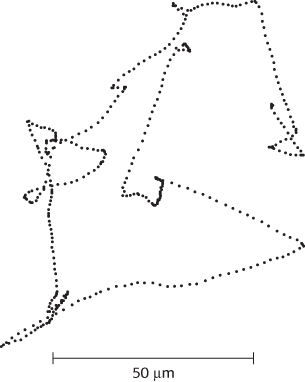
Many species of bacteria respond to changes in their environments by adjusting their swimming behavior. Examination of the paths taken is highly revealing (Figure 35.30). The bacteria swim in one direction for some length of time (typically about a second), tumble briefly, and then set off in a new direction. The tumbling is caused by a brief reversal in the direction of the flagellar motor. When the flagella rotate counterclockwise, the helical filaments form a coherent bundle favored by the intrinsic shape of each filament, and the bacterium swims smoothly. When the rotation reverses, the bundle flies apart because the screw sense of the helical flagella does not match the direction of rotation. Each flagellum then pulls in a different direction and the cell tumbles.
In the presence of a gradient of certain substances such as glucose, bacteria swim preferentially toward the direction of the higher concentration of the substance. Such compounds are referred to as chemoattractants. Bacteria also swim preferentially away from potentially harmful compounds such as phenol, a chemorepellant. The process of moving in specific directions in response to environmental cues is called chemotaxis. In the presence of a gradient of a chemoattractant, bacteria swim for longer periods of time without tumbling when moving toward higher concentrations of the chemoattractant. In contrast, they tumble more frequently when moving toward lower concentrations of the chemoattractant. This behavior is reversed for chemorepellants. The result of these actions is a biased random walk that facilitates net motion toward conditions more favorable to the bacterium.
Chemotaxis depends on a signaling pathway that terminates at the flagellar motor. The signaling pathway begins with the binding of molecules to receptors in the plasma membrane (Figure 35.31). In their unoccupied forms, these receptors initiate a pathway leading eventually to the phosphorylation of a specific aspartate residue on a soluble protein called CheY. In its phosphorylated form, CheY binds to the base on the flagellar motor. When bound to phosphorylated CheY, the flagellar motor rotates in a clockwise rather than a counterclockwise direction, causing tumbling.
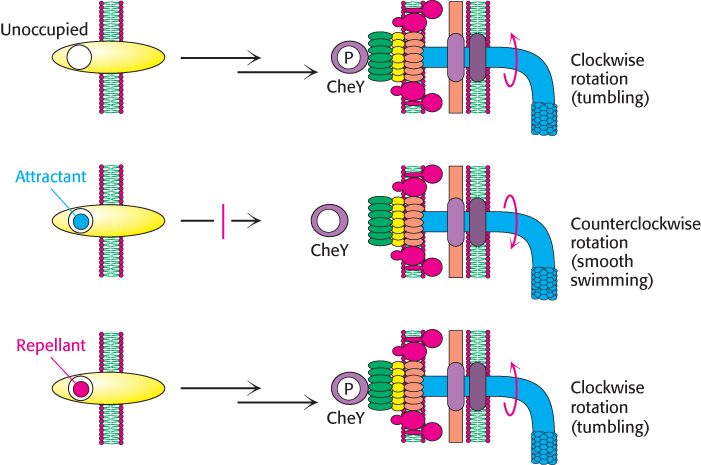
The binding of a chemoattractant to a surface receptor blocks the signaling pathway leading to CheY phosphorylation. Phosphorylated CheY spontaneously hydrolyzes and releases its phosphoryl group in a process accelerated by another protein, CheZ. The concentration of phosphorylated CheY drops, and the flagella are less likely to rotate in a clockwise direction. Under these conditions, bacteria swim smoothly without tumbling. Thus, the reversible rotary flagellar motor and a phosphorylation-
1029
Bacteria sense spatial gradients of chemoattractants by measurements separated in time. A bacterium sets off in a random direction and, if the concentration of the chemoattractant has increased after the bacterium has been swimming for a period of time, the likelihood of tumbling decreases and the bacterium continues in roughly the same direction. If the concentration has decreased, the tumbling frequency increases and the bacterium tests other random directions. The success of this mechanism once again reveals the power of evolutionary problem solving: many possible solutions are tried at random, and those that are beneficial are selected and exploited.